Research
Being alive is ridiculously complicated at both an existential and molecular level. To address the former, I try to simplify the later through physical modeling and, when possible, experimental dissection. Below is a brief summary of a number of my active and past research projects that capture the breadth of my interests.
How do cells strike a balance between metabolism and translation?

Effective coordination of cellular processes is critical to ensure the competitive growth of microbial organisms. Pivotal to this coordination is the appropriate partitioning of cellular resources between protein synthesis via translation and the metabolism needed to sustain it. But how can a cell – which is not much more than a watery bag of proteins – make such a decision? The thrust of my postdoctoral research is figuring out how. To do so, I’ve recently derived and systematically probed a coarse-grained and organism-agnostic theory of microbial growth centered on the dynamic regulation of this resource partitioning. At the core of this regulation is an optimization of the outputs of metabolism and protein translation, mechanistically achieved via the perception of charged- and uncharged-tRNA turnover. We’ve recently shown that this theory is wildy predictive, allowing us to make quantitative descriptions of how things grow in a wide array of conditions, making it an ideal physiological framework to interrogate the dynamics of growth, competition, and adaptation in complex and ever-changing environments.
Using simple math to understand complex proteomes
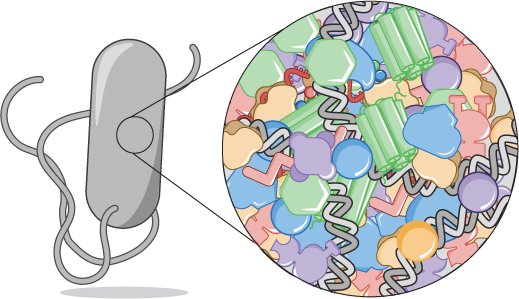
Recent years have seen an experimental deluge interrogating the relationship between bacterial growth rate, cell size, and protein content, quantifying the abundance of proteins across growth conditions with unprecedented resolution. However, we still lack a rigorous understanding of what sets the scale of these quantities and when protein abundances should (or should not) depend on growth rate. Using street-fighting mathematics, I attempt to make order-of-magnitude estimates for the proteinaceous cellular requirements across myriad conditions. These processes include, for example, the requirements for nutrient transport, lipid synthesis, and all steps of the central dogma. Given the wealth of data in our hands for E. coli, we can compare our estimates to actual observations buried in “-omics” measurements. These simple estimates are surprisingly accurate and illustrate the preeminent importance of protein synthesis in defining growth.
Janus-faced molecules and the regulation of genes
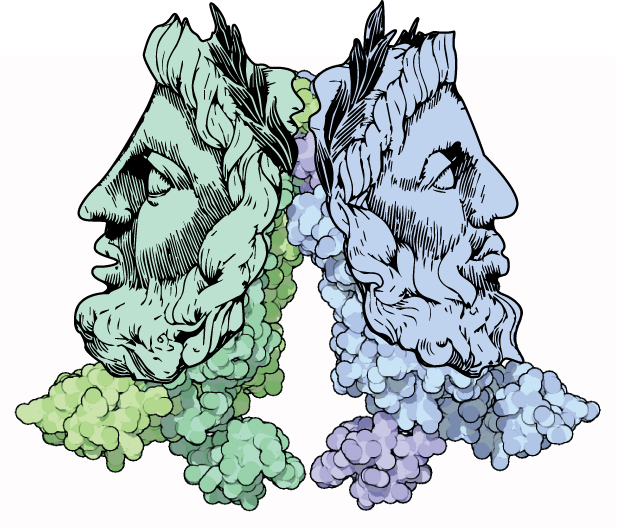
Just like us, molecules can be deceptive. In some conformations, they tightly hold onto DNA with an iron fist, silencing the genes they grasp. In others, they weakly attach to DNA with a velvet glove, allowing the genes to sing. This molecular control is manifest through allosteric regulation – a regulatory mechanism that its discoverer Jacques Monod called “the second secret of life.” My PhD research used a combination of statistical physics, molecular biology, and Bayesian inference to declassify this secret and used it to understand how cells make decisions without any brains.
Quantifying the Anthropocene
The greatest experiment of the last 10,000 years is the presence and action of modern human beings on planet Earth. At this point, the con- sequences of this experiment are being felt on many fronts. Yet, many people still hold the view that because the world is so “huge”, humans cannot really make a substantial impact. In this ongoing research we present a collection of what we have come to view as essential numbers that summarize the broad reach of human action across the planet, presenting a view of the impact of human presence on Earth. These numbers include recent estimates/measurements of the volume of meltwater released from ice-sheets on an annual basis, the year change in ocean acidity from the absorption of CO2, the background plutonium isotope reactivity found in soils stemming from nuclear weapons testing in the 1960’s, to the number of livestock on the planet to give a few of many examples. In collecting and scrutinizing these data, I developed and curate the Human Impacts Database, a internet database similar in spirit to the BioNumbers website that we hope will be used by scientists and the general public alike.